fig1
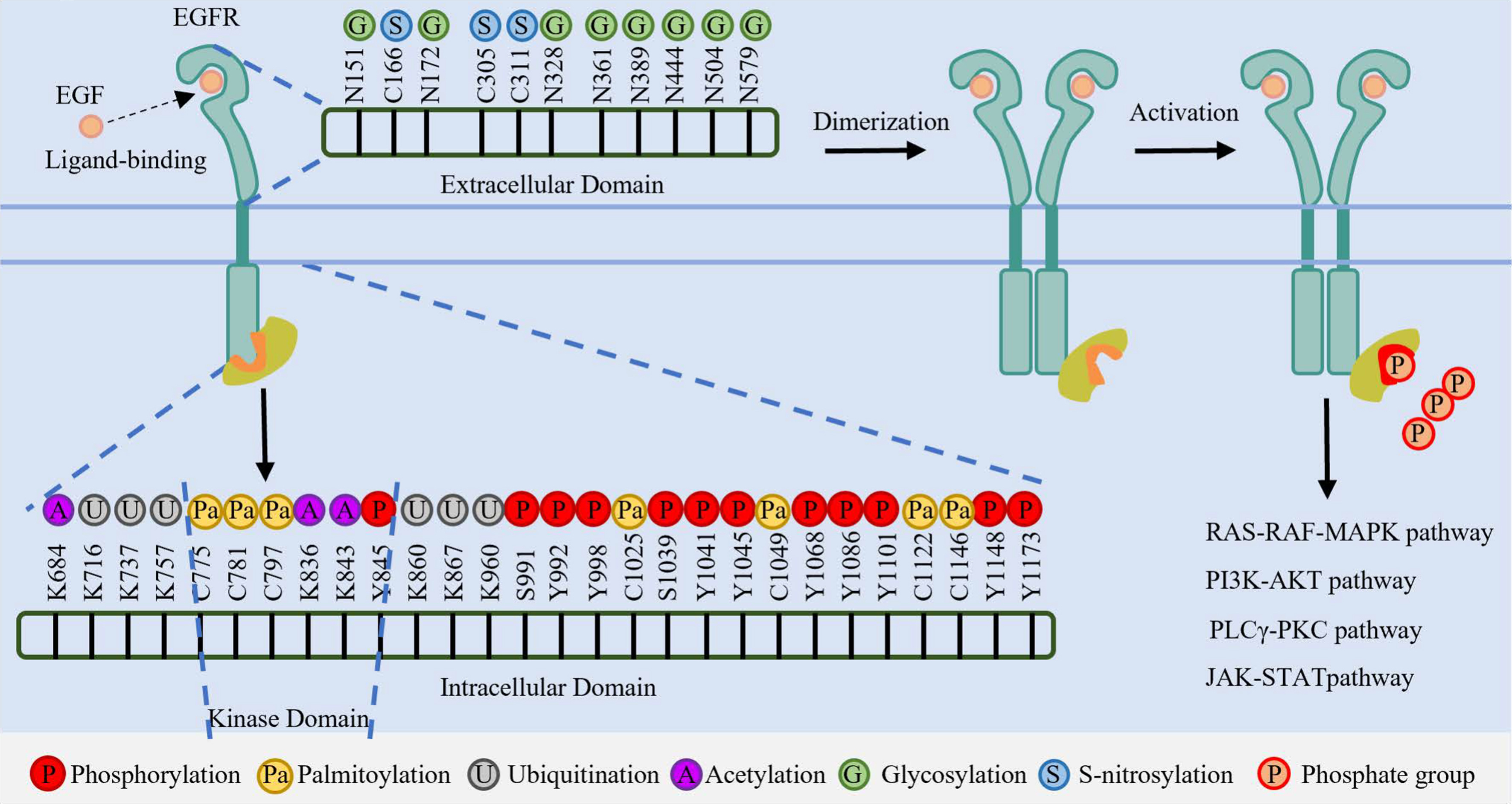
Figure 1. Schematic representation of PTMs on the EGFR extracellular and intracellular domains. Key modification sites, including Glycosylation (N-linked), Acetylation (K-linked), Ubiquitination (K-linked), Phosphorylation (Y-residues), Palmitoylation (C-residues), and S-nitrosylation (C-residues), are indicated. PTMs: Post-translational modifications; EGFR: epidermal growth factor receptor; N: asparagine; K: lysine; Y: tyrosine; S: serine; C: cysteine; EGF: epidermal growth factor; ATP: adenosine triphosphate; RAS: rat sarcoma viral oncogene homolog; RAF: rapidly accelerated fibrosarcoma; MAPK: mitogen-activated protein kinase; PI3K: phosphoinositide 3-kinase; AKT: protein kinase B; PLCγ: phospholipase C gamma; PKC: protein kinase C; JAK: Janus kinase; STAT: signal transducer and activator of transcription.