fig1
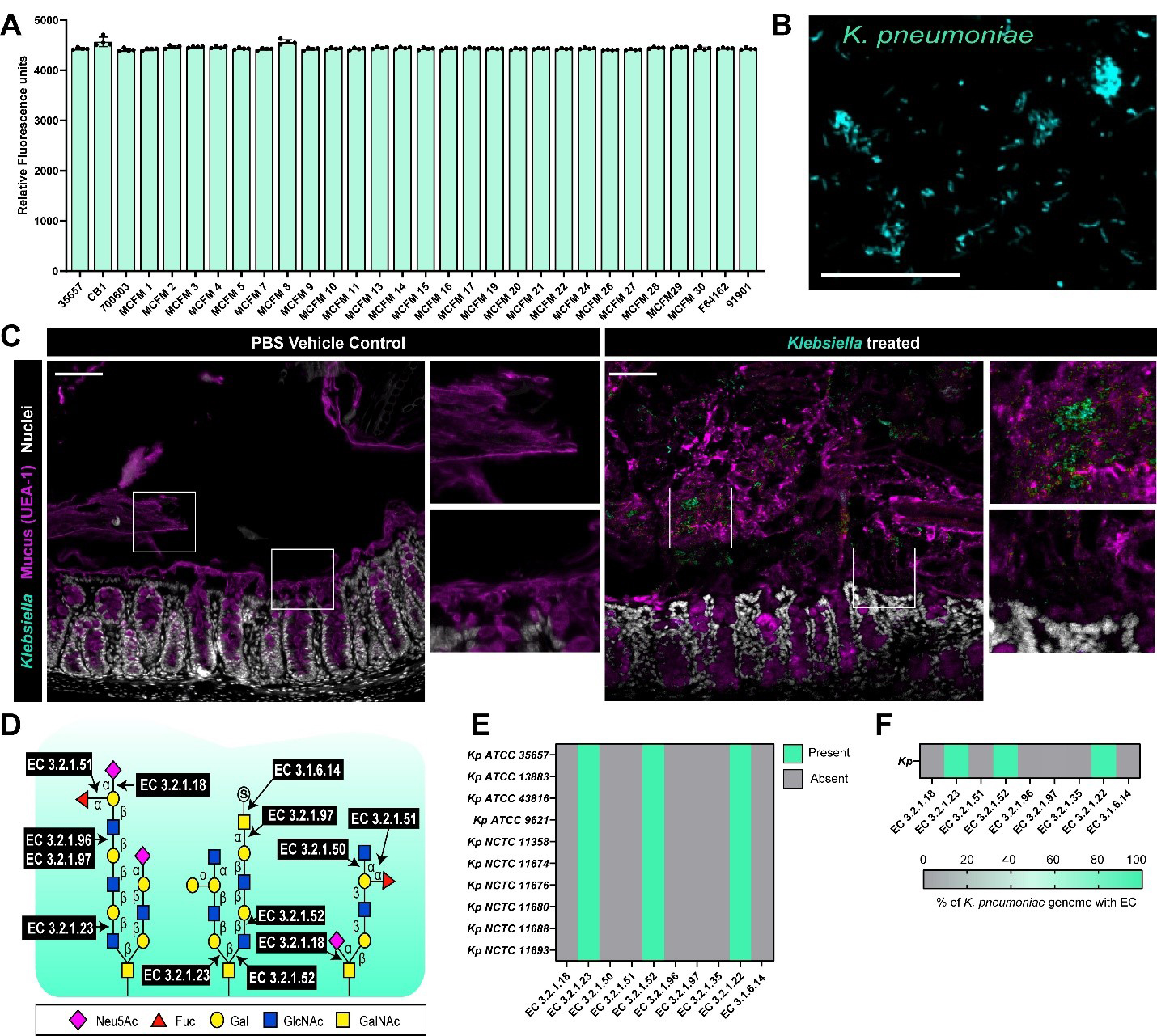
Figure 1. K. pneumoniae can adhere to intestinal mucus. (A) K. pneumoniae strains were fluorescently tagged and added to plates coated with 1 mg/mL dialyzed porcine intestinal mucus and adhesion was examined by a plate reader after 1 h incubation. Data presented as the fluorescence of adhered bacteria to mucus (mean ± standard deviation); (B) Representative fluorescent microscopy image of K. pneumoniae ATCC 35657 adhered to human MUC2-coated coverslips, scale bar = 50 μm; (C) Mice were orally administered with PBS (vehicle control) or K. pneumoniae ATCC 35657 and colons were harvested five days later. K. pneumoniae in the mucus layer was visualized with fluorescent in-situ hybridization (teal), mucus was visualized with UEA-1 staining (purple) and nuclei were identified with Hoechst (white). Scale bar = 50 μm; (D) Schematic of the enzymes required to cleave the bonds attaching mucus-associated sugars; (E) Genome analysis of 10 K. pneumoniae genomes in the IMG database for genes involved in mucus glycan degradation; (F) Genome analysis of the percentage of total K. pneumoniae genomes in the IMG database for genes involved in mucus glycan degradation. Data represents the percentage of genomes with a given EC. Data represents the presence or absence of the target gene. Graphs generated with Graphpad Prism. K. pneumoniae: Klebsiella pneumoniae; PBS: phosphate-buffered saline; EC: Enzyme Commission; ATCC: American Type Culture Collection; MUC2: Mucin 2; IMG: Integrated Microbial Genomes; UEA-1: Ulex europaeus agglutinin I.